Found 2 results
Article
18 February 2025Are Memory Updating Tasks Valid Working Memory Measures? A Meta-Analysis
The memory updating (MU) process is a core component of working memory (WM). To systematically examine the validity of two commonly used MU tasks as WM measures, the present meta-analysis (76 studies, total N = 16,184) synthesized results on the correlation between the two MU tasks and two criterion tasks (working memory capacity (WMC) and fluid intelligence (Gf)). Results indicated a moderate correlation between running memory (RM) and WMC (r = 0.42, 95% CI = [0.37, 0.48]), a weak correlation between n-back and WMC (r = 0.23, 95% CI = [0.19, 0.28]), and moderate correlations between both RM (r = 0.40, 95% CI = [0.35, 0.46]) and n-back (r = 0.34, 95% CI = [0.32, 0.37]) and Gf. Subgroup analyses showed that memory load moderated the correlation between RM and WMC, and stimulus-onset asynchrony moderated the correlation between n-back and both WMC and Gf. The recollection and recognition nature of RM and n-back contributed to their different correlation with WMC, and the involvement of controlled attention in both tasks accounted for their association with Gf. The present meta-analysis indicated that RM is a more valid WM measure in behavioral studies on individual differences.

Article
04 November 2024Diversity and Meta-Analysis of Microbial Differential Abundance in Nasal Metatranscriptomic Profiles of Asthma
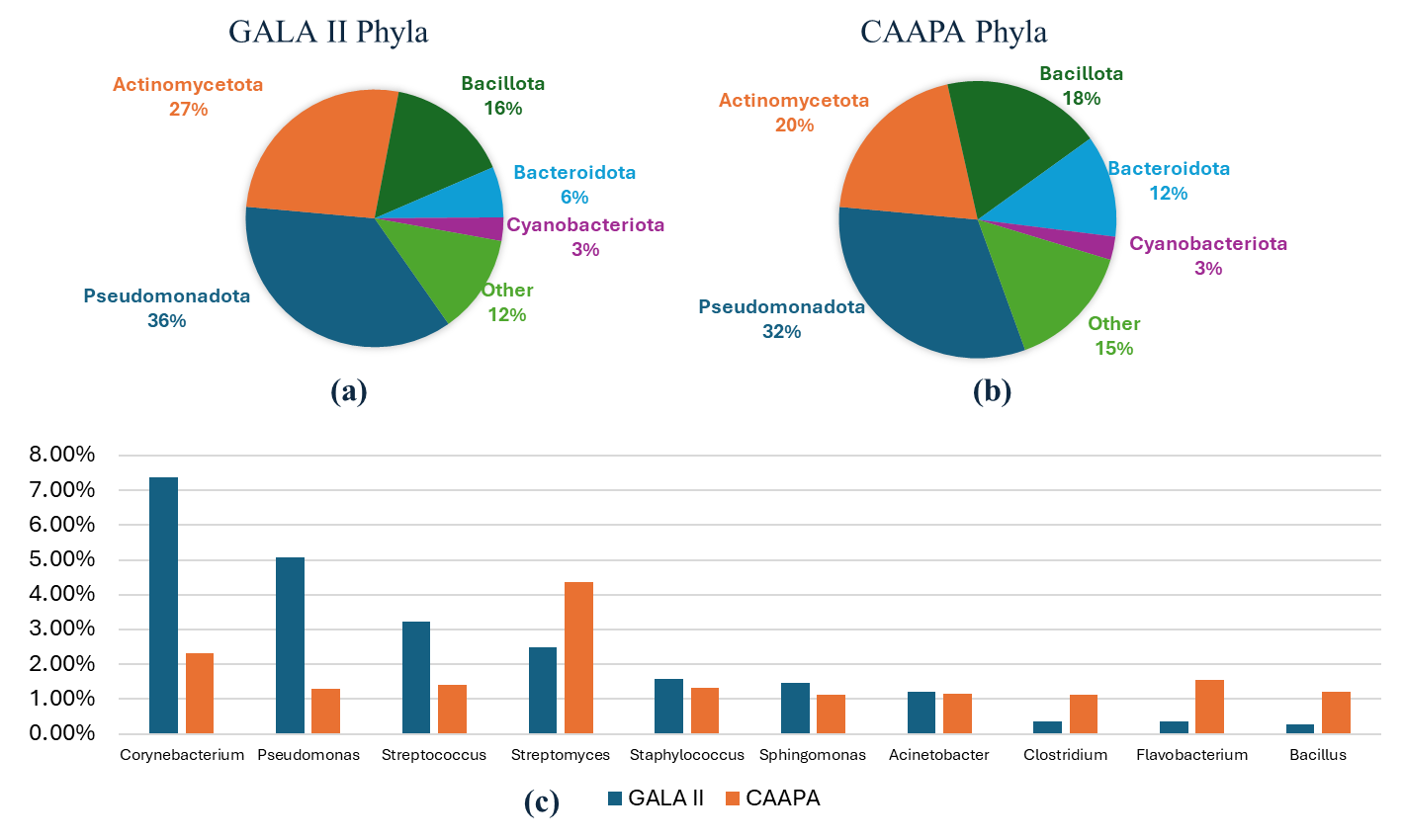
Asthma affects millions worldwide and involves complex genetic, immunological, and environmental factors. The nasal microbiome is increasingly recognized for its role in asthma development, but inconsistent results and small sample sizes have limited a clear understanding. We aimed to clarify the nasal microbiome’s role in asthma using large datasets and meta-transcriptomic analysis. RNA-seq data was analyzed from two large public studies: GALA II (694 children of Puerto Rican heritage; 441 asthmatics, 253 controls) and CAAPA (562 individuals of African ancestry; 265 asthmatics, 297 controls). After quality control and host read removal, microbial reads were annotated using Kraken2. α and β diversity analyses compared microbial diversity between asthmatic and control groups. Differential abundance analysis was conducted separately, controlling for age and sex, with results combined via meta-analysis. We found that asthmatic patients exhibited significantly higher α diversity indices (Shannon, Berger-Parker, Inverse Simpson, Fisher’s) in nasal microbiota compared to controls in GALA II, with similar trends in CAAPA. β diversity analysis showed significant differences in microbial composition in GALA II data. Differential abundance analysis identified 20 species in GALA II and 9 species in CAAPA significantly associated with asthma. Meta-analysis revealed 11 species significantly associated with asthma, including Mycobacterium_tuberculosis. Our study demonstrates increased nasal microbiome α diversity in asthmatic patients and identifies specific microbial species associated with asthma risk. These findings enhance understanding of asthma pathogenesis from the nasal microbiome perspective and may inform future research and therapeutic strategies.